Scottish Crop Map 2019
The ‘Scottish Crop Map’ shows all the agricultural fields in Scotland categorised into the likely main crop types which were grown in 2019.
Methodology
The methods presented here provide an overview of the steps taken to produce crop predictions in Scotland in 2019. The methods are broken down into pre-processing, processing, and modelling steps. To explore the methods further please visit the GitHub repository or email the team.
Product Overview of the Scottish Crop Map
The Crop Map of Scotland is a polygon vector dataset containing a subset of crop types in Scotland. The dataset contains 385,028 fields classifying Scotland into 4 main crop types, grassland, or unknown crop types (where no prediction was made).
Non-agricultural land is excluded from our analysis. Permanent grassland is included in the analysis as it contributes to a large proportion of agricultural fields. Crop types with a smaller amount of fields, including temporary grassland and crops grown in polytunnels, are excluded from the analysis due to not enough ground truth data being available to train the random forest model on.
A supervised classification algorithm (Random Forest Model) was used on data acquired from Sentinel-1 radar images during the period March to October 2019. The results were quality-assured against ground truth data (using Rural Payments and Inspections Division (RPID) inspection data) as well as summary data provided by the Scottish Agricultural Census Team and external stakeholders.
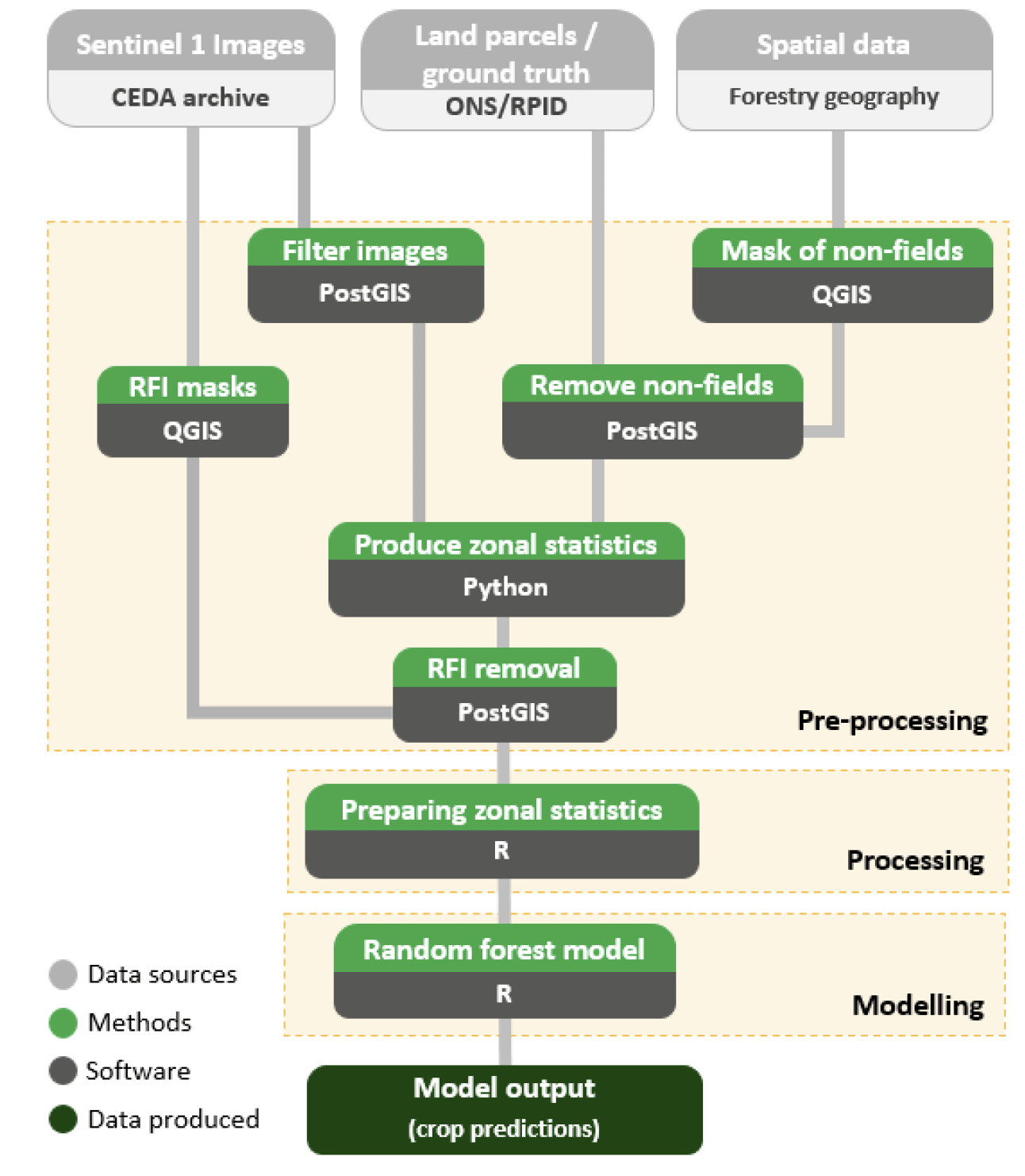
Pre-processing
Pre-processing steps were required to clean and combine data sets ready for analysis. Figure 1 shows the steps which were undertaken in this project. JNCC produced and supplied us with analysis-ready Sentinel-1 radar images. These were accessed through the CEDA archive in JNCC's virtual machine (JASMIN).
Mask of non-fields
Spatial data was collected and used to create a mask of non-field areas in QGIS software. A maximum field size threshold was also applied to remove very large areas likely to be used for rough grazing and a spatial dataset was produced of the mask area. A minimum field size threshold (an area under 1,300m2) was also applied to avoid the use of small areas which did not produce viable zonal statistics.
Remove non-fields
The mask identifying non-field areas, and extremely large or small fields, was then applied to the land parcels data set (in the software PostGIS) resulting in only land parcels we identified as fields being taken forward.
RFI masks
Areas suffering from Radio Frequency Interference (RFI) were manually identified in QGIS and masks were produced. A lookup containing field and date combinations for these RFIs was also created.
Filter Sentinel-1 images
Sentinel-1 images provided to us by JNCC were filtered to only those covering our particular selection of Scottish fields. This was done in PostGIS.
Produce zonal statistics
Python scripts were written (functioning within JNCC's JASMIN environment) to access these analysis-ready Sentinel-1 radar images within the CEDA archive (covering only Scotland). Once retrieved, radar images and known field boundaries (excluding extremely large and small fields) were used to summarise pixel values within each field into the mean values of the radar backscatter (for VH and VV polarisation).These values are referred to as the 'zonal statistics' for each field.
RFI removal
Fields containing RFIs were filtered out once the zonal statistics were produced using a lookup file of date-field combinations which had RFIs in them. Some areas were seriously affected by radar interference – this included the Dumfries area where we were unable to collect enough satellite data to use within the model, resulting in these fields being unclassified.
Processing
Preparing zonal statistics
Zonal statistics (which now excluded RFI areas) were averaged to form six-day blocks (rather than near-daily observations) to consolidate temporally close points into one variable. Other processing steps were performed to prepare the data for use within the random forest model. This was done in R.
Modelling
Random forest model
The zonal statistics were used to train a random forest model on only those fields with known crop types (ground truth data).
Once the zonal statistics were read into R, interpolation was carried out. This dealt with the remaining missing data so that more fields could be predicted by the model. Subsets were created to contain known fields (that have been assigned a crop type) and all (known and unknown) fields.
Some data preparation was conducted on the known fields dataset – any fields that still contained missing values (after averaging into six-day blocks and interpolating) alongside non-crops were removed. Crop types with a small amount of fields (< 25) were excluded from the known dataset and the model. This resulted in the model only including spring barley, winter barley, winter wheat, spring oats and (permanent) grassland.
Afterwards, the known dataset was split into training and test datasets using a 60/40 ratio. This was used to ensure there was enough data for smaller crop types in the test dataset. The training dataset was used to create a random forest model and model improvements were made using accuracies from the test dataset. These were: variable selection, selecting the number of trees (ntree), selecting the number of variables used at each split in a tree (mtry) and identifying the number of training fields to use for each crop (sampsize).
Once the model had been evaluated on the full dataset (containing all fields), a probability cut-off was selected based on the predicted probabilities. A probability cut-off of 0.48 was set for the model which allowed for the best balance between under- and overestimating crop area. For those fields that didn't meet the cut-off, it was assumed the model was less confident and thus were predicted to be NA (missing).
Code
The code used in (a) generating the zonal statistics and (b) running the random forest model are available on GitHub in the following repository:
https://github.com/cropmapteam/Scottish-Crop-Map-2019-Publication
Contact
Email: agric.stats@gov.scot