Coronavirus (COVID-19): modelling the epidemic (issue no. 60)
Latest findings in modelling the COVID-19 epidemic in Scotland, both in terms of the spread of the disease through the population (epidemiological modelling) and of the demands it will place on the system, for example in terms of health care requirement.
Technical Annex
Epidemiology is the study of how diseases spread within populations. One way we do this is using our best understanding of the way the infection is passed on and how it affects people who catch it to create mathematical simulations. Because people who catch Covid-19 have a relatively long period in which they can pass it on to others before they begin to have symptoms, and the majority of people infected with the virus will experience mild symptoms, this "epidemiological modelling" provides insights into the epidemic that cannot easily be measured through testing e.g. of those with symptoms, as it estimates the total number of new daily infections and infectious people, including those who are asymptomatic or have mild symptoms.
Modelling also allows us to make short-term forecasts of what may happen with a degree of uncertainty. These can be used in health care and other planning. The modelling in this research findings is undertaken using different types of data which going forward aims to both model the progress of the epidemic in Scotland and provide early indications of where any changes are taking place.
The delivery of the vaccination programme will offer protection against severe disease and death. The modelling includes assumptions about compliance with restrictions and vaccine take-up. Work is still ongoing to understand how many vaccinated people might still spread the virus if infected. As Covid-19 is a new disease there remain uncertainties associated with vaccine effectiveness. Furthermore, there is a risk that new variants emerge for which immunisation is less effective.
How the modelling compares to the real data as it emerges
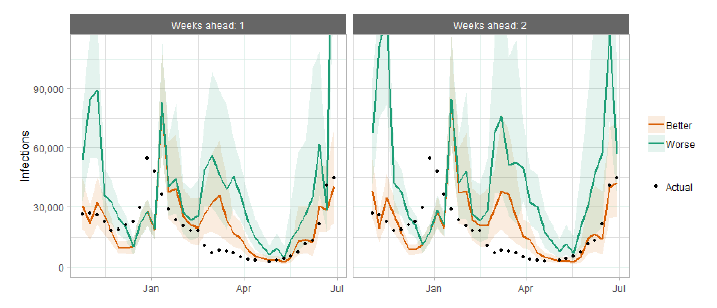
The following charts show the history of our modelling projections in comparison to estimates of the actual data. The infections projections were largely accurate during October to mid-December and from mid‑January onward. During mid-December to mid-January, the projections underestimated the number of infections, due to the unforeseen effects of the new variant.
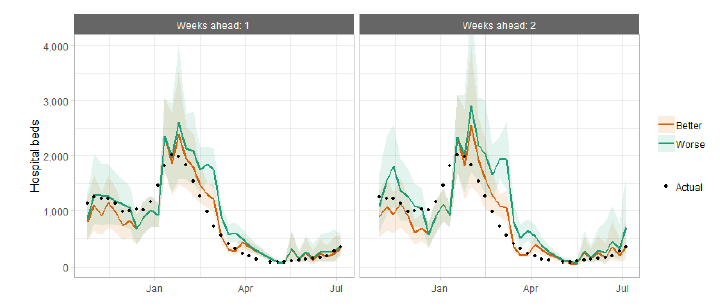
Hospital bed projections have generally been more precise than infections estimates due to being partially based on already known information about numbers of current infections, and number of people already in hospital. The projections are for number of people in hospital due to Covid-19, which is slightly different to the actuals, which are number of people in hospital within 28 days of a positive Covid-19 test.
As with hospital beds, ICU bed projections have generally been more precise than infections. The projections are for number of people in ICU due to Covid-19. The actuals are number of people in ICU within 28 days of a positive Covid-19 test up to 20 January, after which they include people in ICU over the 28 day limit.

How outliers are identified in the wastewater data
On occasion, samples of wastewater (WW) produce extremely high measurements of viral RNA concentration that are unlikely to be representative of actual Covid-19 prevalence and revert to low levels subsequently – even in samples taken very soon afterwards. These account for approximately 2% of samples.
These spikes usually remain unexplained. Possible explanations could include an imperfect sampling process, incidents of dumping of virus contaminated waste, or lab contamination. Sometimes, though, there can be a reasonable explanation for the spike, such as an outbreak centred on a hospital or care home, or an influx of holiday-makers.
Identification of such spikes improve interpretability of data visualisations, show clearer underlying trends in aggregate levels, avoid spurious alerts, and improve the reliability of measures of uncertainty. It is especially desirable to be able to detect anomalies immediately after the WW RNA value is recorded, since this is when decisions may need to be made, and new samples might be obtained to recheck Covid-19 levels.
Biomathematics and Statistics Scotland (BioSS) has designed an automatic procedure for detecting these spikes, based on modelling their degree of divergence from case trends and prior WW RNA trends.
After first normalising WW RNA values with respect to flow, the outlier removal methodology has these steps:
1. For each WW observation, a measure of its effectiveness in predicting subsequent Covid-19 increases is calculated, based on its ratio with local case rates in the week immediately after.
2. The observation's ratio with prior case rates is calculated, as well a measure of how much the rate of change in WW RNA levels compares to previous changes in RNA at the same site.
3. The value in step 1 is modelled as a function of the two calculated values in step 2, using a generalised additive model with appropriately chosen logarithmic transformations.
4. For each newly observed WW measurement, the same covariates as in step 2 can be calculated, and the model we fitted may be used to forecast the new measurement's predictive effectiveness, calculating a propensity score for that value to be an outlier between 0 and 10. When a decision is required, a threshold then may be imposed on this propensity using a preliminary analysis with manually identified spikes.
An example of the output from the algorithm is show below in Figure 22.
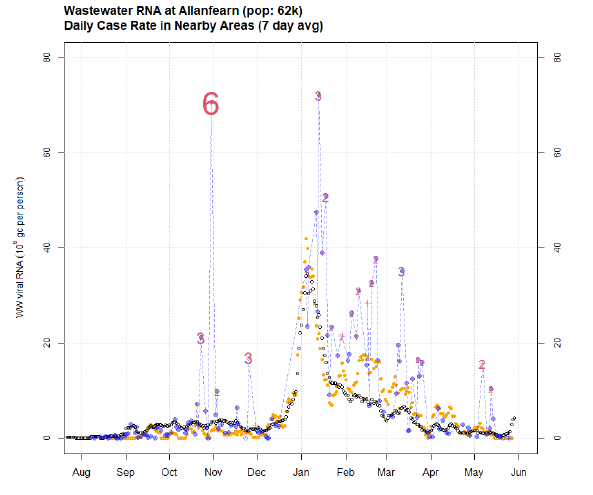
Comparing to manually chosen outliers, a threshold of 4 picks up about 70% of manually detected spikes and misclassifies around 0.7% of non-anomalous values. Note that this threshold choice is quite conservative, choosing to avoid classifying borderline cases as outliers. Applying this threshold in the example above means only the late October spike is identified as an outlier.
Use of outlier identification
For aggregate values like national or local authority means, anomalous values are removed to produce more reliable results, though sometimes this may leave aggregates entirely missing major cities in an area. We are considering further how to improve the management of these cases.
However, it is generally believed that a positive WW value, even an erroneously large one, nevertheless indicates the presence of Covid-19 in a region. This means that outliers should not be removed when it comes to presence/absence analyses.
The outlier detection methodology is currently implemented in graphical visualisations in the weekly WW national reporting, for the tables of local authority averages, and in interval-based summaries of Covid-19 levels for local authorities (Table 2).
How is wastewater data used in our modelling?
The Scottish Government has historically used either deaths or cases, as published by Public Health Scotland (PHS), to inform its model to estimate current R values, incidence figures and growth rates.
In recent months, these research findings have explained how an estimate of cases can be made by examining the levels of Covid-19 RNA in wastewater, collected throughout Scotland and adjusted for population and local changes in intake flow rate.
We have developed our modelling such that it is possible to calculate the main nowcast outputs by using this wastewater data, instead of the case data from PHS.
The Scottish WW data is population weighted averages for normalised Wastewater Covid levels. The units are provided in 1 million gene copies per person per day, which roughly matches with cases per 100,000 per day. This is converted into daily cases at a national level. The model makes an allowance for the proportion of infections which are positively identified as cases (using a comparison with the ONS Covid Infection Survey[14]), and then uses a Bayesian method to estimate the key variables throughout the pandemic.
We are currently only using the wastewater data for Scottish cases, but are working with colleagues in the other UK nations to use their wastewater data in a similar way.
| Probability of exceeding (cases per 100k) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Local Authority (LA) | 20 | 50 | 100 | 150 | 300 | 500 | 750 | 1000 | 2000 |
| Aberdeen City | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 5-15% | 0-5% |
| Aberdeenshire | 75-100% | 75-100% | 75-100% | 75-100% | 25-50% | 15-25% | 5-15% | 5-15% | 0-5% |
| Angus | 75-100% | 75-100% | 75-100% | 75-100% | 75-100% | 25-50% | 15-25% | 5-15% | 0-5% |
| Argyll and Bute | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 5-15% | 0-5% | 0-5% | 0-5% |
| City of Edinburgh | 75-100% | 75-100% | 75-100% | 75-100% | 75-100% | 25-50% | 25-50% | 25-50% | 15-25% |
| Clackmannanshire | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 5-15% | 0-5% | 0-5% | 0-5% |
| Dumfries & Galloway | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 15-25% | 5-15% | 0-5% | 0-5% |
| Dundee City | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 50-75% | 25-50% | 25-50% | 15-25% |
| East Ayrshire | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 15-25% | 15-25% | 0-5% |
| East Dunbartonshire | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 5-15% | 0-5% |
| East Lothian | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 5-15% | 0-5% |
| East Renfrewshire | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 5-15% | 0-5% |
| Falkirk | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 5-15% | 0-5% |
| Fife | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 15-25% | 5-15% |
| Glasgow City | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 25-50% | 25-50% | 15-25% |
| Highland | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 5-15% | 0-5% | 0-5% | 0-5% |
| Inverclyde | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 5-15% | 0-5% | 0-5% |
| Midlothian | 75-100% | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 15-25% | 5-15% | 0-5% |
| Moray | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 5-15% | 5-15% | 5-15% | 0-5% |
| Na h-Eileanan Siar | 25-50% | 5-15% | 0-5% | 0-5% | 0-5% | 0-5% | 0-5% | 0-5% | 0-5% |
| North Ayrshire | 75-100% | 75-100% | 75-100% | 75-100% | 25-50% | 25-50% | 15-25% | 5-15% | 0-5% |
| North Lanarkshire | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 25-50% | 15-25% | 5-15% |
| Orkney Islands | 25-50% | 25-50% | 15-25% | 5-15% | 0-5% | 0-5% | 0-5% | 0-5% | 0-5% |
| Perth and Kinross | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 15-25% | 15-25% | 5-15% | 0-5% |
| Renfrewshire | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 15-25% | 15-25% | 5-15% |
| Scottish Borders | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 5-15% | 5-15% | 0-5% | 0-5% |
| Shetland Islands | 25-50% | 5-15% | 0-5% | 0-5% | 0-5% | 0-5% | 0-5% | 0-5% | 0-5% |
| South Ayrshire | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 25-50% | 15-25% | 5-15% | 0-5% |
| South Lanarkshire | 75-100% | 75-100% | 75-100% | 75-100% | 25-50% | 25-50% | 25-50% | 15-25% | 15-25% |
| Stirling | 75-100% | 75-100% | 75-100% | 75-100% | 25-50% | 15-25% | 5-15% | 0-5% | 0-5% |
| West Dunbartonshire | 75-100% | 75-100% | 75-100% | 75-100% | 50-75% | 25-50% | 5-15% | 5-15% | 0-5% |
| West Lothian | 75-100% | 75-100% | 75-100% | 75-100% | 75-100% | 25-50% | 25-50% | 15-25% | 5-15% |
What levels of Covid-19 are indicated by wastewater (WW) data?
Table 2 provides population weighted daily averages for normalised WW Covid-19 levels in the weeks beginning the 26th June and 3rd July, with no estimate for error. This is given in Million gene copies per person, which approximately corresponds to new cases per 100,000 per day. Coverage is given as percentage of LA inhabitants covered by a wastewater Covid‑19 sampling site delivering data during this period[15].
| Local authority (LA) | Average daily WW case estimate, with outliers included |
Average daily WW case estimate, with outliers removed |
Coverage[16] | ||
|---|---|---|---|---|---|
| w/b 26th June | w/b 3rd July | w/b 26th June | w/b 3rd July | ||
| Aberdeen City | 87.0 | 136.0 | 87.0 | 136.0 | 80% |
| Aberdeenshire | 32.0 | 62.0 | 32.0 | 57.0 | 52% |
| Angus | 85.0 | 201.0 | 85.0 | 201.0 | 56% |
| Argyll and Bute | 0.0 | 10.0 | 0.0 | 10.0 | 18% |
| City of Edinburgh | 172.0 | 94.0 | 172.0 | 94.0 | 96% |
| Clackmannanshire | 33.0 | 46.0 | 33.0 | 46.0 | 92% |
| Dumfries & Galloway | 25.0 | 67.0 | 25.0 | 28.0 | 32% |
| Dundee City | 106.0 | 248.0 | 106.0 | 248.0 | 100% |
| East Ayrshire | 30.0 | 57.0 | 30.0 | 54.0 | 69% |
| East Dunbartonshire | 83.0 | 114.0 | 83.0 | 114.0 | 99% |
| East Lothian | 154.0 | 99.0 | 154.0 | 99.0 | 65% |
| East Renfrewshire | 62.0 | 73.0 | 62.0 | 73.0 | 95% |
| Falkirk | 67.0 | 81.0 | 67.0 | 81.0 | 69% |
| Fife | 73.0 | 86.0 | 73.0 | 86.0 | 80% |
| Glasgow City | 70.0 | 97.0 | 70.0 | 97.0 | 98% |
| Highland | 32.0 | 79.0 | 32.0 | 78.0 | 33% |
| Inverclyde | 40.0 | 60.0 | 40.0 | 60.0 | 92% |
| Midlothian | 161.0 | 108.0 | 161.0 | 108.0 | 88% |
| Moray | 21.0 | 32.0 | 7.0 | 32.0 | 70% |
| Na h-Eileanan Siar | 0.0 | 3.0 | 0.0 | 3.0 | 21% |
| North Ayrshire | 23.0 | 22.0 | 23.0 | 22.0 | 93% |
| North Lanarkshire | 37.0 | 126.0 | 37.0 | 126.0 | 95% |
| Orkney Islands | 4.0 | 6.0 | 4.0 | 6.0 | 34% |
| Perth and Kinross | 32.0 | 35.0 | 32.0 | 35.0 | 45% |
| Renfrewshire | 44.0 | 89.0 | 44.0 | 89.0 | 57% |
| Scottish Borders | 12.0 | 21.0 | 10.0 | 21.0 | 38% |
| Shetland Islands | 0.0 | 0.0 | 0.0 | 0.0 | 29% |
| South Ayrshire | 23.0 | 27.0 | 23.0 | 27.0 | 82% |
| South Lanarkshire | 58.0 | 70.0 | 58.0 | 65.0 | 84% |
| Stirling | 28.0 | 44.0 | 28.0 | 44.0 | 63% |
| West Dunbartonshire | 11.0 | 76.0 | 11.0 | 76.0 | 98% |
| West Lothian | 50.0 | 86.0 | 46.0 | 87.0 | 85% |