Coronavirus (COVID-19): modelling the epidemic (issue no.81)
Latest findings in modelling the COVID-19 epidemic in Scotland, both in terms of the spread of the disease through the population (epidemiological modelling) and of the demands it will place on the system, for example in terms of health care requirement.
Technical Annex
Epidemiology is the study of how diseases spread within populations. One way we do this is using our best understanding of the way the infection is passed on and how it affects people who catch it to create mathematical simulations. Because people who catch Covid-19 have a relatively long period in which they can pass it on to others before they begin to have symptoms, and the majority of people infected with the virus will experience mild symptoms, this "epidemiological modelling" provides insights into the epidemic that cannot easily be measured through testing e.g. of those with symptoms, as it estimates the total number of new daily infections and infectious people, including those who are asymptomatic or have mild symptoms.
Modelling also allows us to make short-term forecasts of what may happen with a degree of uncertainty. These can be used in health care and other planning. The modelling in this research findings is undertaken using different types of data which going forward aims to both model the progress of the epidemic in Scotland and provide early indications of where any changes are taking place.
The delivery of the vaccination programme will offer protection against severe disease and death. The modelling includes assumptions about compliance with restrictions and vaccine take-up. Work is still ongoing to understand how many vaccinated people might still spread the virus if infected. As Covid-19 is a new disease there remain uncertainties associated with vaccine effectiveness. Furthermore, there is a risk that new variants emerge for which immunisation is less effective.
How the modelling compares to the real data as it emerges
The method of producing the medium term projections (figures 5 - 7) uses the published actual numbers of infections, hospital admissions and ICU admissions directly, rather than modelling them from the beginning of the epidemic. This means the projections begin from the point the published data ends.
There is no prediction interval around the actual infections in Figure 5 because there is no longer any uncertainty from simulating infections during this period. There is still uncertainty in the ascertainment rate, which is represented by the whiskers around the actual infections.
The prediction intervals around the actual hospital and ICU occupancy in Figures 6 and 7 now represent uncertainty in the assumptions for the hospitalisation rate and hospital length of stay, rather than uncertainty in the number of infections. These confidence intervals are created by applying sensitivity analysis to the assumptions.
The following charts show the history of our modelling projections in comparison to estimates of the actual data. The infections projections were largely accurate during October to mid-December 2020 and from mid‑January 2021 onwards. During mid-December 2020 to mid‑January 2021, the projections underestimated the number of infections, due to the unforeseen effects of the new variant.
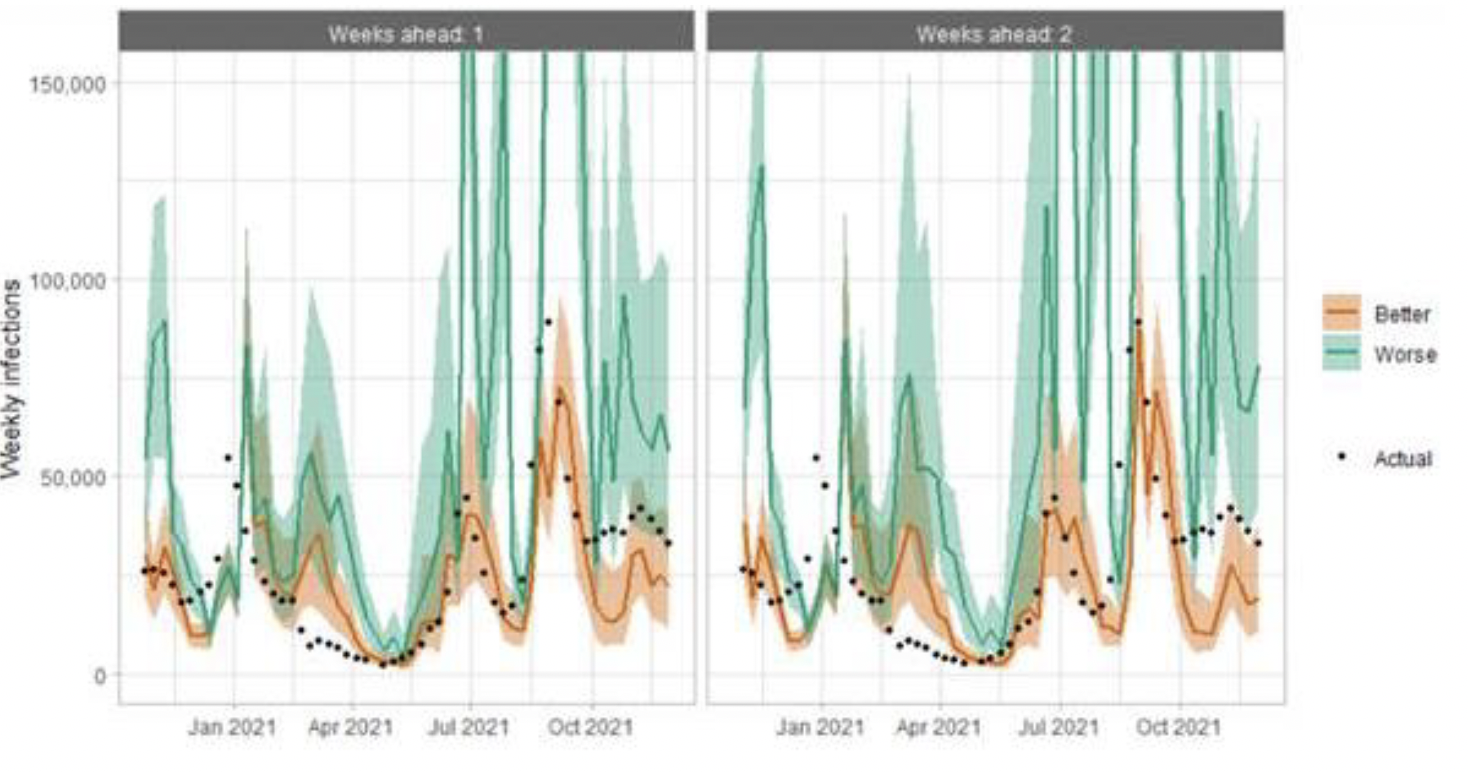
Hospital bed projections have generally been more precise than infections estimates due to being partially based on already known information about numbers of current infections, and number of people already in hospital. The projections are for number of people in hospital due to Covid-19, which is slightly different to the actuals, which are number of people in hospital within 28 days of a positive Covid-19 test.
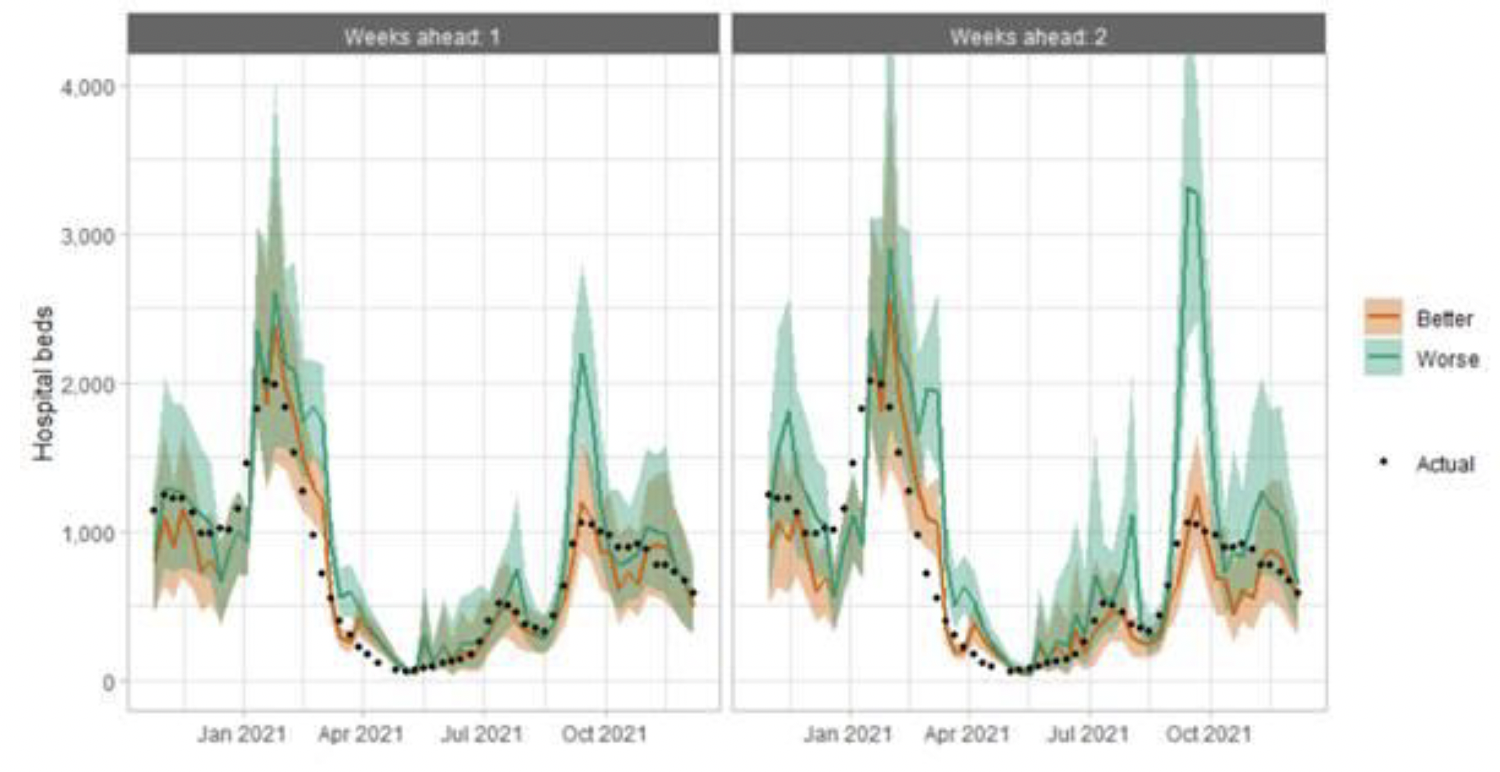
As with hospital beds, ICU bed projections have generally been more precise than infections. The projections are for number of people in ICU due to Covid-19. The actuals are number of people in ICU within 28 days of a positive Covid-19 test up to 20 January 2021, after which they include people in ICU over the 28 day limit.
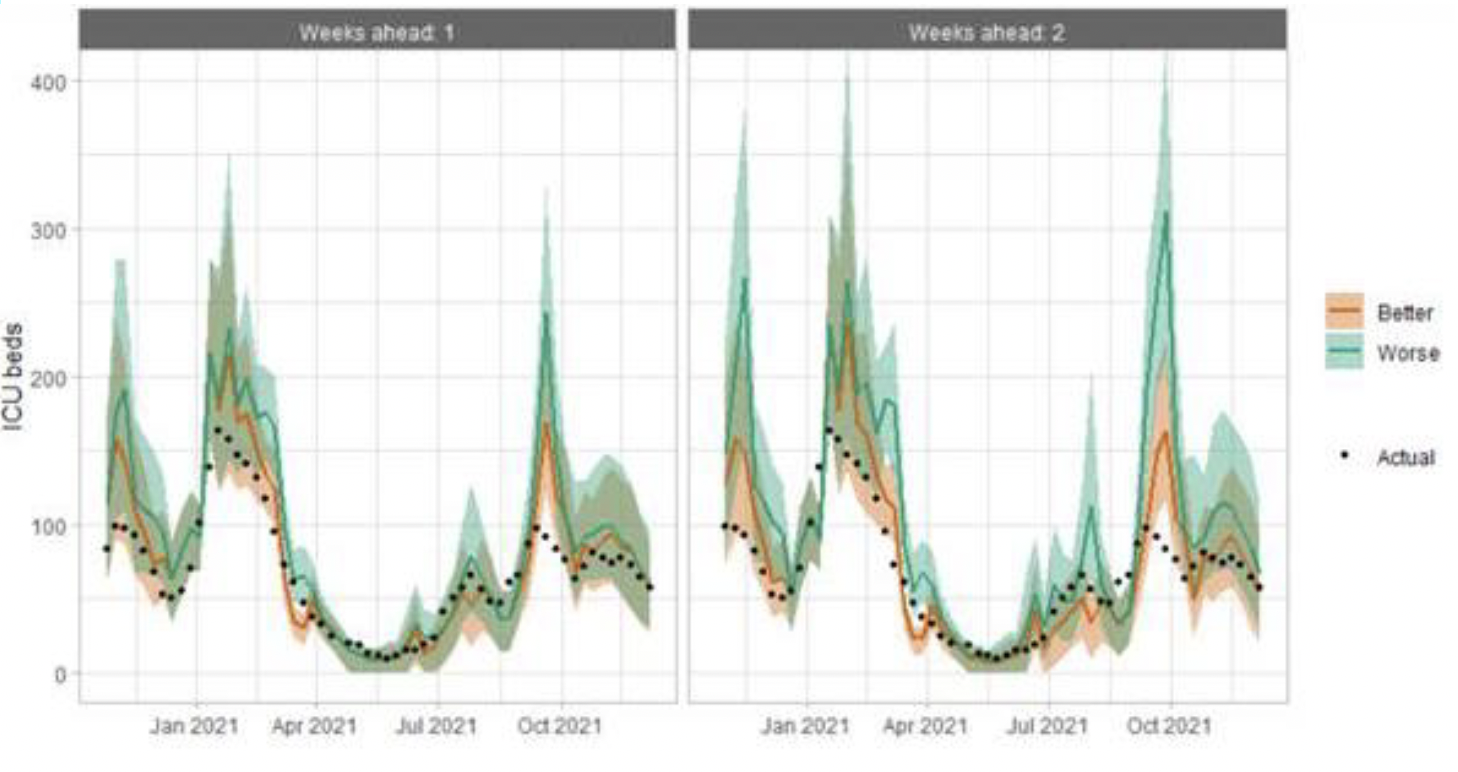
How is the doubling time for the Omicron variant calculated
In a recent WHO report[15] it was confirmed that in PCR tests one of the three target genes of the SARS-CoV-2 virus is not detected (called S‑gene target failure, SGTF). This test can therefore be used as an indicator for the Omicron variant.
The number of total S-gene tests and the proportion which test positive for SGTF in Scotland are provided daily by PHS. Based on a methodology developed by academics at the University of Manchester, a statistical model is based on some underlying parameters of the Omicron variant. This is fitted against the actual number of S-gene test and the proportion of these which are positive for SGTF using maximum likelihood. This allows for a best estimate of the underlying parameters, including the relative growth advantage of the Omicron variant, which leads to an estimate of the doubling time of the variant.
There remains significant unknowns about the Omicron variant, which makes it difficult to estimate projections with any degree of confidence. Key to this is any measure of severity, transmissibility and vaccine escape. This information will be incorporated into the various models as and when it becomes known.
Which local authorities are likely to experience high levels of Covid-19 in two weeks' time
| Probability of exceeding (cases per 100K) | ||||
|---|---|---|---|---|
| Local Authority (LA) | 50 | 100 | 300 | 500 |
| Aberdeen City | 75-100% | 75-100% | 15-25% | 0-5% |
| Aberdeenshire | 75-100% | 75-100% | 5-15% | 0-5% |
| Angus | 75-100% | 75-100% | 25-50% | 0-5% |
| Argyll and Bute | 75-100% | 75-100% | 15-25% | 0-5% |
| City of Edinburgh | 75-100% | 75-100% | 50-75% | 15-25% |
| Clackmannanshire | 75-100% | 75-100% | 50-75% | 15-25% |
| Dumfries & Galloway | 75-100% | 75-100% | 50-75% | 5-15% |
| Dundee City | 75-100% | 75-100% | 15-25% | 0-5% |
| East Ayrshire | 75-100% | 75-100% | 75-100% | 50-75% |
| East Dunbartonshire | 75-100% | 75-100% | 75-100% | 25-50% |
| East Lothian | 75-100% | 75-100% | 50-75% | 15-25% |
| East Renfrewshire | 75-100% | 75-100% | 50-75% | 15-25% |
| Falkirk | 75-100% | 75-100% | 75-100% | 25-50% |
| Fife | 75-100% | 75-100% | 50-75% | 15-25% |
| Glasgow City | 75-100% | 75-100% | 50-75% | 25-50% |
| Highland | 75-100% | 75-100% | 5-15% | 0-5% |
| Inverclyde | 75-100% | 75-100% | 25-50% | 15-25% |
| Midlothian | 75-100% | 75-100% | 25-50% | 15-25% |
| Moray | 75-100% | 75-100% | 25-50% | 5-15% |
| Na h-Eileanan Siar | 75-100% | 50-75% | 15-25% | 0-5% |
| North Ayrshire | 75-100% | 75-100% | 25-50% | 5-15% |
| North Lanarkshire | 75-100% | 75-100% | 50-75% | 25-50% |
| Orkney Islands | 50-75% | 25-50% | 0-5% | 0-5% |
| Perth and Kinross | 75-100% | 75-100% | 25-50% | 0-5% |
| Renfrewshire | 75-100% | 75-100% | 25-50% | 15-25% |
| Scottish Borders | 75-100% | 75-100% | 25-50% | 0-5% |
| Shetland Islands | 25-50% | 15-25% | 0-5% | 0-5% |
| South Ayrshire | 75-100% | 75-100% | 50-75% | 25-50% |
| South Lanarkshire | 75-100% | 75-100% | 50-75% | 25-50% |
| Stirling | 75-100% | 75-100% | 25-50% | 5-15% |
| West Dunbartonshire | 75-100% | 75-100% | 50-75% | 15-25% |
| West Lothian | 75-100% | 75-100% | 50-75% | 25-50% |
What levels of Covid-19 are indicated by wastewater data?
Table 2 provides population weighted daily averages for normalised WW Covid-19 levels in the weeks beginning 24th November and 1st December 2021, with no estimate for error. This is given in Million gene copies per person, which approximately corresponds to new cases per 100,000 per day. Coverage is given as percentage of LA inhabitants covered by a wastewater Covid‑19 sampling site delivering data during this period[16].
| Local authority (LA) | w/b 24th November | w/b 1st December | Coverage |
|---|---|---|---|
| Aberdeen City | 99 | 53 | 80% |
| Aberdeenshire | 76 | 58 | 29% |
| Angus | 81 | 64 | 55% |
| Argyll and Bute | 35 | 44 | 12% |
| City of Edinburgh | 85 | 76 | 98% |
| Clackmannanshire | 83 | – | 0% |
| Dumfries & Galloway | 67 | – | 8% |
| Dundee City | 89 | 64 | 100% |
| East Ayrshire | 49 | 88 | 69% |
| East Dunbartonshire | 94 | 102 | 99% |
| East Lothian | 106 | 82 | 74% |
| East Renfrewshire | 76 | 57 | 89% |
| Falkirk | 91 | 133 | 69% |
| Fife | 68 | 120 | 77% |
| Glasgow City | 86 | 79 | 71% |
| Highland | 72 | 60 | 48% |
| Inverclyde | 57 | 53 | 98% |
| Midlothian | 82 | 76 | 73% |
| Moray | 115 | 89 | 14% |
| Na h-Eileanan Siar | 4 | – | 0% |
| North Ayrshire | 67 | 66 | 92% |
| North Lanarkshire | 73 | 103 | 33% |
| Orkney Islands | 26 | 18 | 34% |
| Perth and Kinross | 76 | 86 | 38% |
| Renfrewshire | 54 | 66 | 97% |
| Scottish Borders | 84 | 40 | 27% |
| Shetland Islands | – | – | 0% |
| South Ayrshire | 54 | 94 | 88% |
| South Lanarkshire | 66 | 61 | 58% |
| Stirling | 54 | 35 | 53% |
| West Dunbartonshire | 70 | 80 | 98% |
| West Lothian | 89 | 136 | 95% |